This page was last revised for version 5.0.10.
Curation Assistant
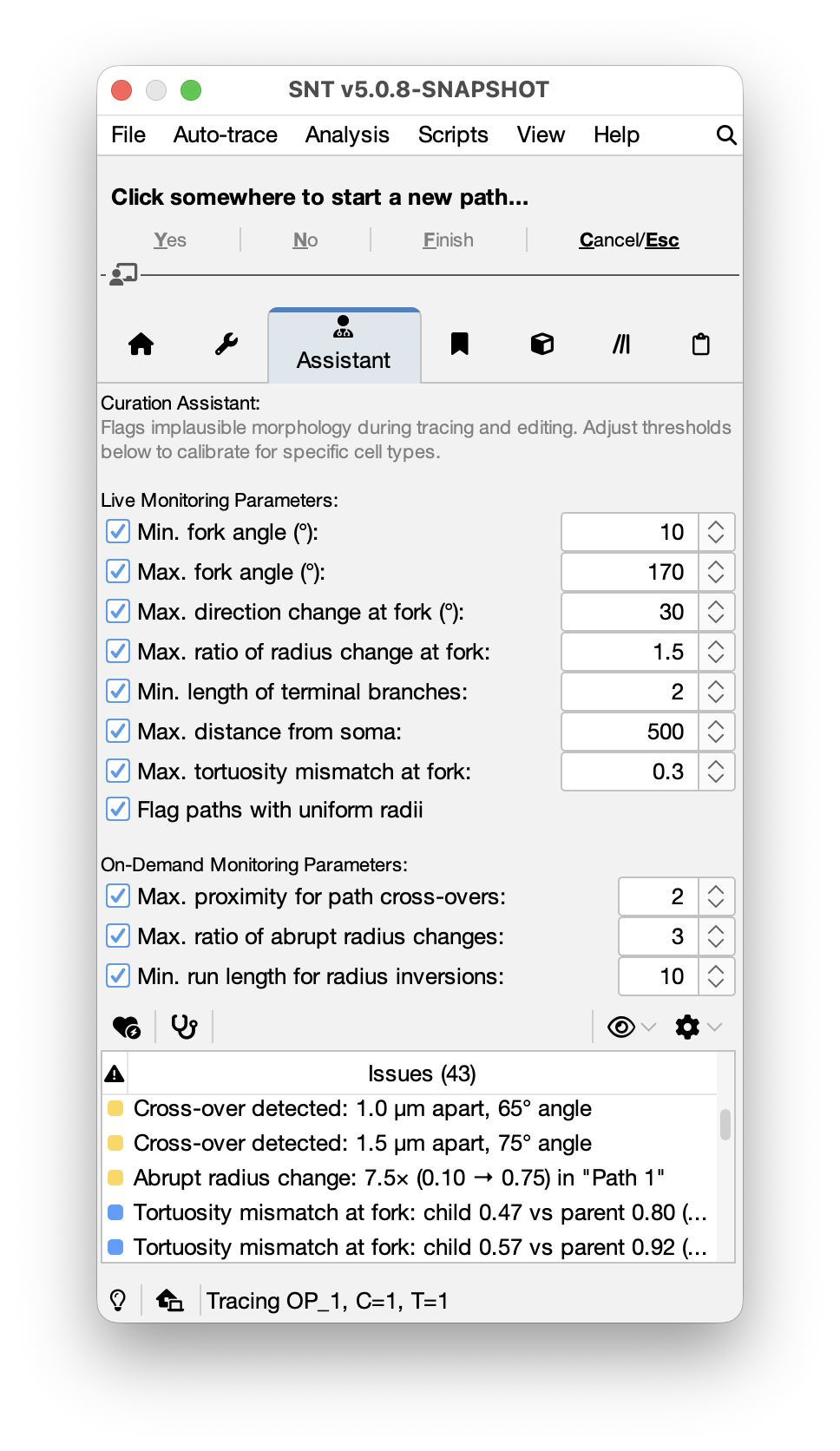
The Curation Assistant is a tab in the main dialog that monitors the morphological plausibility of neuron reconstructions in real time (see Curation vs Proofreading). It flags suspect tracing operations as they happen, e.g., near-parallel fork angles, abrupt radius changes, U-turns at branch points, and so on. It also provides an on-demand “full scan” that detects whole-reconstruction issues such as path overlaps and radius inversions. All parameters can be adjusted in the GUI, saved to reusable preset files, or calibrated automatically from reference reconstructions. The Assistant is organized into four sections:
- Live Monitoring Parameters: Lightweight checks that run inline during interactive tracing (at fork initiation, segment completion, path finalization, and node editing)
- On-Demand Monitoring Parameters: More Computationally-intensive checks that scan the full reconstruction when explicitly triggered
- Toolbar: Action buttons for toggling live monitoring, running a full scan, filtering warnings by severity, and an options menu for calibration and presets
- Warnings Table: Displays current issues sorted by severity. Double-click a row to navigate to the warning location in the image
Live Monitoring Parameters
These checks execute in sub-millisecond time and are evaluated automatically during tracing when live monitoring is enabled. Each check has a checkbox to enable or disable it and a field (where applicable) to set its threshold
Min. fork angle (°)
Warns when the angle between a parent path and a child branch is suspiciously narrow, suggesting the child runs nearly parallel to the parent. The angle is computed from the parent’s direction at the branch point and the child’s initial heading (first 5 nodes). A fork angle below this threshold produces a warning. Range: 0–90°
Max. fork angle (°)
Warns when the fork angle is too wide, as when the child branch nearly doubles back on the parent, which is anatomically rare and often indicates a tracing error. A fork angle above this threshold produces a warning. Range: 90–180°
Min. fork angle and Max. fork angle control a single underlying check (Branch angle). The check is enabled whenever at least one of the two checkboxes is selected.
Max. direction change at fork (°)
Detects potential U-turns where a child path reverses the parent’s trajectory. The check compares the parent’s tangent vector at the branch point with the child path’s initial direction vector (first 5 nodes) using a dot-product angle. When the angle between the two vectors falls below this threshold, it suggests the child is continuing backward along the parent rather than branching away. Range: 0–90°
Max. ratio of radius change at fork
Flags forks where the child neurite’s caliber differs sharply from the parent’s at the branch point. The parent radius is sampled at the branch node; the child radius is the median of its first 5 nodes. The ratio \(\frac{child\,radius}{parent\,radius}\) is compared against this threshold. Range: 1.0–10.0
Min. length of terminal branches
Catches stub branches that are too short to be anatomically meaningful, in case they were generated by accidental clicks or tracing artifacts. Only terminal paths (with no children) are evaluated. A terminal branch whose length (in image units) falls below this threshold is flagged. Range: 0.1–100.0 (in image units)
Max. distance from soma
Flags primary paths whose starting node is unexpectedly far from any soma marker. When soma nodes are present, the Euclidean distance (in image units) from the path’s first node to the nearest soma is measured. Range: 10.0–10000.0 (in image units)
Max. tortuosity mismatch at fork
Warns when a child path has a markedly different sinuosity from its parent, measured by contraction. The absolute difference in contraction between parent and child is compared against this threshold. Both paths must have at least 5 nodes for the check to apply. Range: 0.05–1.0
Flag paths with uniform radii
Identifies paths where every node shares the same radius value, which typically means radii remain at a default constant. This check has no threshold field: it is either enabled or disabled.
On-Demand Monitoring Parameters
These checks scan the full set of paths in the reconstruction. They are triggered by clicking the Run Full Scan button.
Max. proximity for path cross-overs
Detects crossovers: regions where topologically unconnected paths run suspiciously close to each other, suggesting duplicate or overlapping tracings.
Internally uses CrossoverFinder with a proximity radius. Self-crossovers and direct parent-child pairs are excluded.
Range: 0.1–100.0
Max. ratio of abrupt radius changes
Finds adjacent nodes within a single path that have a sudden radius jump. This often results from radius-fitting errors or misaligned cylinder models. The ratio \(\frac{\max(r_1, r_2)}{\min(r_1, r_2)}\) between consecutive nodes is compared against this threshold. Range: 1.5–20.0
Min. run length for radius inversions
Flags sustained centripetal radius increases along a path. Since neurites generally taper as they extend away from the soma, a long run of increasing radii suggests an error. A “run” is a consecutive sequence of nodes where each radius exceeds the previous. When a run’s length meets or exceeds this threshold, a warning is produced. Range: 3–100 (no. of nodes)
Min. signal contrast ratio
Measures how well each path stands out from the background in the image. At every node (excluding the first and last), a perpendicular cross-section is sampled using a Bresenham line that extends 3× the path radius (minimum 5 px) on each side. The sampled intensities are split by percentile: the lower quartile (P25) estimates background, the upper quartile (P75) estimates signal. The contrast ratio is defined as:
\[\frac{\mathrm{median}(\text{signal}) + 1}{\mathrm{median}(\text{background}) + 1}\]A path whose median contrast ratio falls below this threshold is flagged. The +1 offset prevents division by zero and stabilizes the ratio for dim images. When set to -1 (Auto), the threshold is computed automatically from the image’s intensity range as \(\frac{I_{\max} + 1}{I_{\min} + 1} \times \frac{1}{2}\), representing half of the best achievable contrast ratio for the image. The computed value is displayed in the spinner after the scan completes; use the undo button to reset to auto mode. This check requires an image to be loaded and image statistics to be computed.
Range: -1 (auto) or 1.0–100.0
Toolbar Actions
Live Monitoring Toggle
Enables or disables real-time plausibility checking during interactive tracing. When enabled, the assistant monitors into fork initiation, segment completion, path finalization, and node editing events, running all enabled live checks automatically.
Run Full Scan
Triggers a comprehensive scan of all paths in the current reconstruction using both live and on-demand parameters. Live checks are applied to every parent–child fork relationship; on-demand checks scan the path collection as a whole. Results are displayed in the warnings table.
Filter by Severity
A dropdown with checkboxes for each severity level (Errors, Warnings, Informational Notes) that controls which type of issues appear in the table.
Options Menu
Auto-tuning
Calibrate Thresholds from Traced Cells… Opens a file chooser for selecting reconstruction files. The selected cells are loaded, and all thresholds are inferred from their morphological statistics using percentile analysis (5th percentile for lower-bound thresholds, 95th percentile for upper-bound). The calibrated values are applied to the panel immediately. See Advanced Curation Tasks section for details on the underlying algorithm.
Built-in Presets
Lists curation presets bundled with SNT. These are calibrated from NeuroMorpho reference data and provide sensible starting points for common cell types. Selecting a preset loads its thresholds and enabled states into the panel. Presets include:
| Preset | Cell type | Species | Publication(s) | NeuroMorpho archive |
|---|---|---|---|---|
| CA1 Dendrites | CA1 pyramidal (basal + apical dendrites) | Mouse | Mellström et al., 2016 | DeFelipe |
| CA3 Dendrites | CA3 pyramidal (basal + apical dendrites) | Rat | Ishizuka et al., 1995 | Amaral |
| Eurydendroid Cells | Eurydendroid | Larval zebrafish | Kunst et al., 2019 | Baier |
| Kenyon Cells | Kenyon | Adult Drosophila | Zheng et al., 2018 | Bock |
| Martinotti Cells | Martinotti interneuron (axons/dendrites) | Mouse | Goldberg et al., 2004, Trevelyan et al., 2006, Nikolenko et al., 2007 | Yuste |
| Purkinje Dendrites | Purkinje neuron (dendrites) | Mouse | Chen et al., 2013 | Dusart |
| RGC Dendrites | Retinal ganglion cell (dendrites) | Mouse | Werginz et al., 2020 | Fried |
User Presets
Lists preset files (.curation extension) found in the curations/ subdirectory of the SNT workspace (typically ~/SNT_workspace/curations/). This section of the menu includes:
- Create From Current Parameters… Saves the current thresholds and enabled states to a new
.curationfile (described here). - Reload User Presets: Rescans the curations directory and refreshes the menu
- Reveal User Presets Directory: Opens the curations directory in the system file manager
Warnings Table
The table displays warnings sorted by severity (Errors first, then Warnings, then Informational Notes). Each row shows a color-coded severity indicator and a message describing the issue.
Double-click a row to navigate to the warning’s spatial location in the image. The view centers on the affected node and zooms to the configured visiting zoom level.
Right-click the table for additional options: clearing all warnings, setting the visiting zoom level, resetting zoom to the current image magnification, and detaching/docking the table into a separate floating window.
Seed Review
The Curation Assistant also accepts warnings contributed by external programs: Warnings produced outside SNT can be imported as Warning seeds, and loaded into the assistant. These entries:
- Carry the check name “Seed Review” so they’re easier to spot (and filter) among the algorithmically-generated warnings
- Use the seed’s 3D position as the warning location, so the standard double-click-to-navigate behavior flies the canvas to the seed.
- Their severity level (Info / Warning / Error) is manually chosen: Use whichever level matches the review intent
- Encode the seed’s confidence, type, and source in the message
A typical workflow:
- Import seeds into the Seeded Tracing Assistant (e.g., a CSV file of predicted problematic locations generated by a deep-learning pipeline)
- Adjust the confidence range and sort the table so the candidates of interest are at the top
- Select the rows to be reviewed, right-click, and choose Send Selected Seeds to Curation Assistant… → As Warning (or As Info Message / As Error, as appropriate)
- The action will import the selected seeds and activate to the Assistant tab so that you can step through the new entries via double-click navigation
NB: Imported seeds are not de-duplicated: sending the same selection twice produces two rows
Detecting Crossovers
A crossover is a spot where at least two neurites pass very close to each other in space (so they may look like they intersect in the image) but they are not connected in the reconstructed graph (i.e., there is no shared node / true topological join). Identification of crossover sites is thus useful to disambiguate overlaps between neurites and spot possible tracing mistakes, such as missed branch-points or false merges.

Overview of a crossover between two neurites.
Left: Seem from top, the two neurites seem to intersect. Right: rotation to front view reveals that the two paths are juxtaposed in the XZ plane.
Algorithm
Crossover events between two paths, Path A and Path B, are detected as follows:
- Seeds:
All tracing nodes from both paths plus segment midpoints are used as seed points (midpoints help catch “T‑like” geometries where a node lies near the middle of another path’s segment). Midpoints are flagged so they can be treated specially later - Proximity mining:
A uniform 3‑D grid is built with cell size equal to the proximity radius. For each seed, the 27‑cell neighborhood (the seed’s cell ± 1 in x/y/z) is queried; candidate pairs are kept if their Euclidean distance ≤ proximity and they satisfy a series of optional criteria - Candidate grouping:
Candidate pairs are grouped by unordered (Path A, Path B) pairs, then sorted by index (iA, iB) and deduplicated. Each group is split into monotonic “runs”. A run is accepted if its length is within a specified cutoff, or if it is a single‑pair run that touches an endpoint - Geometric verification:
For each accepted run, all corresponding segment pairs are examined: the closest points and distances between segments are computed, as well as an orientation‑invariant approach angle (0–90°) from local tangents. The center of the crossover event is the mean of closest‑point midpoints; the median distance and median angle summarize the run. Optional angle thresholds can be applied - Merging:
Nearby events (centers within proximity) are merged: the center is averaged, participants are unioned, index windows are merged, the distance becomes the minimum of medians, and the angle becomes the mean of medians - Validation:
A final post‑hoc filter keeps an event only if at least one path node from a participant path is near the event center. This removes spurious “floating” events
Obtaining Crossover Locations
From the GUI, the easiest way to list crossover events is to use the Bookmark option in the Navigator Toolbar. Alternatively, running the curation assistant’s full scan will also run the detection routine. For advanced detections, scripting is advised. In a script, detection settings are specified in a Config object, example:
// groovy
import sc.fiji.snt.util.CrossoverFinder
cfg = new CrossoverFinder.Config()
.proximity(2.0) // The Neighborhood radius [real‑world units (e.g., µm)] used 1) for the coarse grid query during candidate mining, and 2) to merge nearby events
.thetaMinDeg(0.0) // Minimum approach angle (0–90°) required to accept an event. Use 10–20° to suppress nearly parallel neurites
.minRunNodes(2) // Minimum length of a “near‑pair run”. Neurites need to have at list this no. of nodes at the crossover site for it to be detected
.sameCTOnly(true) // Only compare paths with the same channel and frame?
.includeSelfCrossovers(false) // Allow detections within the same neurite? Generally keep false
.includeDirectChildren(false) // Allow detections within a path and its direct child? Generally keep false
.nodeWitnessRadius(-1.0) // Post‑hoc filter: A crossover event is only kept if at least one participant path has an actual node (not a midpoint) within this radius of the event center. Default (-1) instructs proximity valueOnce the config is defined, events can be detected from any collection of paths:
// groovy
import sc.fiji.snt.Tree
tree = Tree.fromFile("path/to/a/swc/file.swc")
paths = tree.list()
events = CrossoverFinder.find(paths, cfg)
for (ev in events) {
println("x=%.2fµm, y=%.2fµm, z=%.2fµm, angle=%.1f°, d=%.2fµm, paths=%d%n",
ev.x, ev.y, ev.z, ev.medianAngleDeg, ev.medianMinDist, ev.participants.size())
xyzct = ev.xyzct() // pixel-space + avg C/T
// Optional: create a bookmark or navigate to xyzct[0..2]
}Advanced Curation Tasks
Running Checks Programmatically
All curation classes live in the sc.fiji.snt.analysis.curation package. The three main entry points are:
PlausibilityCheckThe checks themselvesPlausibilityMonitorThe runtime coordinator, managing both tiers of checksPlausibilityCalibratorThreshold inference and preset I/O
A typical usage in a script would like this:
import sc.fiji.snt.Tree
import sc.fiji.snt.analysis.curation.PlausibilityMonitor
import sc.fiji.snt.analysis.curation.PlausibilityCheck
// Load a reconstruction
tree = Tree.fromFile("/path/to/cell.swc")
// Create and configure the monitor
monitor = new PlausibilityMonitor()
// Adjust individual check thresholds
ba = monitor.getLiveCheck(PlausibilityCheck.BranchAngle.class)
ba.setMinAngleDeg(15.0)
ba.setMaxAngleDeg(90.0)
rj = monitor.getDeepCheck(PlausibilityCheck.RadiusJumps.class)
rj.setMaxJumpRatio(2.0)
// Run a full scan (live + on-demand checks on all paths)
warnings = monitor.runFullScan(tree.list())
warnings.each { w ->
println("${w.severity()}: ${w.message()}")
}
Calibrating Thresholds Programmatically
From Local Files
PlausibilityCalibrator infers thresholds from reference reconstructions. For each parameter, it collects the relevant metric across all parent–child forks (live checks) or all paths (deep checks) and derives a threshold from percentile statistics.
Lower-bound parameters (e.g., minimum fork angle) use a lower percentile; upper-bound parameters (e.g., maximum radius ratio) use an upper percentile.
import sc.fiji.snt.Tree
import sc.fiji.snt.analysis.curation.PlausibilityCalibrator
import sc.fiji.snt.analysis.curation.PlausibilityMonitor
// Load reference cells
trees = [
Tree.fromFile("/path/to/cell1.swc"),
Tree.fromFile("/path/to/cell2.swc"),
Tree.fromFile("/path/to/cell3.swc"),
]
// Calibrate
calibrator = new PlausibilityCalibrator(trees)
calibrator.setUpperPercentile(95.0) // default
calibrator.setLowerPercentile(5.0) // default
result = calibrator.calibrate()
// Print the summary table: A formatted summary showing each check's
// inferred threshold, sample size, mean, sd, and the percentile used
println(result.toTable())
// Apply to a monitor
monitor = new PlausibilityMonitor()
result.applyTo(monitor)
From NeuroMorpho.org
SNT includes a script template (Tracing › Calibrate Curation From NeuroMorpho…) that downloads reference cells by their NeuroMorpho.org identifiers, calibrates thresholds, and saves a .curation preset.
The template accepts parameters for the preset name, a comment, and the upper/lower percentiles. Edit the cellIds list in the script body with identifiers for your cell type of interest.
Saving and Loading Curation Presets Programatically
Presets can be saved and loaded programmatically:
import sc.fiji.snt.analysis.curation.PlausibilityCalibrator
import sc.fiji.snt.analysis.curation.PlausibilityMonitor
monitor = new PlausibilityMonitor()
// ... configure monitor thresholds ...
// Save to a .curation file
outFile = new File("/path/to/my-preset.curation")
PlausibilityCalibrator.save(monitor, outFile, "My custom preset")
// Load from a .curation file
PlausibilityCalibrator.load(outFile, monitor)
// Load a built-in preset bundled with SNT
PlausibilityCalibrator.loadBuiltIn("dummy1", monitor)
