The content of this page has not been vetted since shifting away from MediaWiki. If you’d like to help, check out the how to help guide!
In addition to the more generic Automatic Loader and Stack loaders, we also offer single-purpose loaders for images acquired on a Zeiss Lightsheet Z.1 or a diSPIM controlled via MicroManager.
Zeiss Lightsheet Z.1 Dataset (Bioformats)
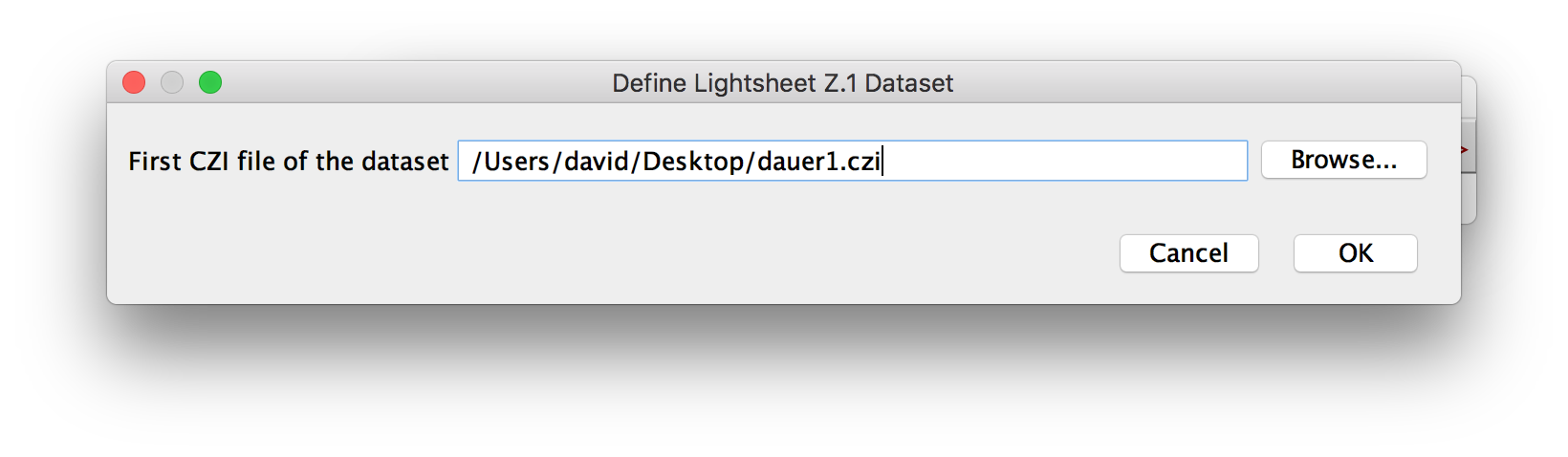
If you select to load a Zeiss Lightsheet Z.1 Dataset, you will first be asked for the first .czi-file in the dataset (Step 1). If your dataset is just a single file, pick that file. If your dataset consists of multiple files, pick the one without a numeric suffix.
We will then parse the dataset and show the metadata we could extract in the next dialog (Step 2). There are a few changes you can make at this moment:
- The Angle, Tiles, Channels, Illumination Directions and Time Points that were detected in the dataset will be shown and you can rename them.
- The calibration (pixel distances) will be shown. You can select Modify calibration to change it manually in the next step.
- The rotation axis of a multi-Angle dataset will be shown at the bottom of the dialog.
- Ticking Modify rotation axis will let you choose the axis manually in the next step.
- Ticking Apply rotation to dataset will transform the individual views according to the rotations from metadata. This should give you a rough alignment to start with.
- Finally, due to a bug in BioFormats, all stacks in a file may be reported to have the same number of z-slices even if their number of slices differ (smaller stacks will be filled with zeroes to reach the size of the largest stack). If you tick Fix Bioformats stack size bug, we will inspect the image data and remove the zero-valued volume at the end of the stack.
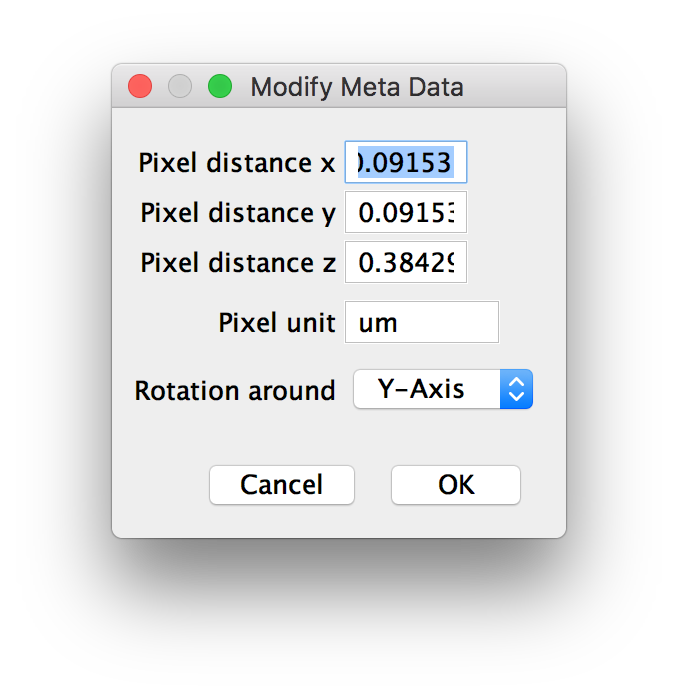
If you chose to modify the calibration or the rotation axis, a third dialog will be displayed in which you can manually set the following thinks (Step 3):
- The pixel distance in every dimension (x,y and z).
- The unit the pixel distance is measured in.
- Which axis the different acquisition angles are rotated around.
After working through these steps, your dataset will be opened in BigStitcher.
MicroManager diSPIM Dataset
TODO
Go back to the dataset definition overview
Go back to the main page