The CMP-BIA tools is a package for ImageJ/Fiji which will perform image segmentation and registration. For a fast integration of our plugins you can use our update site.
All source codes are publicly available as Maven project (see the GitHub repository). The API in this package can be also used for further development of other Java/ImageJ features related to Image Processing.
Note, all included methods are mainly related to medical imaging but it can be also used in the fields.
Please note that these plugins available through Fiji, is based on publications. If you use it successfully for your research please be so kind to cite our related work in References.
Center for Machine Perception (CMP) is a university research center performing fundamental and applied research in computer vision, robotics, machine learning, pattern recognition, and mathematics. Biomedical Imaging Algorithms group (BIA) headed by Jan Kybic develops new algorithms for biomedical image processing.
Plugins
The package currently contains a few plugins:
- jSLIC superpixels - is segmentation method for clustering similar regions - superpixels - in given image which are usually used for other segmentation techniques. The only two parameters are average (initial) size of each superpixel and rigidity parameter in range [0,1].
jSLIC - superpixels
Recently, SLIC (Simple Linear Iterative Clustering) was introduced for general images and presented as a powerful intermediate phase for further image segmentation, classification and registration. SLIC is an adaptation of the k-means algorithm for superpixel generation with two important distinctions: (a) the weighted distance measure combines colour (using the CIELAB colour space, which is widely considered as perceptually uniform for small colour distances) and spatial proximity and (b) the search space is reduced by limiting to a region 2Sx2S, proportional to the superpixel size S. The search space reduction has a great impact on the speed of whole algorithm.
We made a Java-based open source implementation jSLIC - the superpixel clustering with better performance than the original SLIC. Moreover, we proposed a different regularisation parameter, which influences the compactness of resulting superpixels and propose a default value r=0.2. The new post-processing step gives more reliable superpixels shapes, with no need of decreasing superpixel size.
How to use the plugin
jSLIC interface
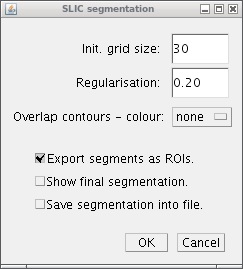
As you can see, the interface to the plugin contains parameters for superpixel configuration and also its final visualisation.
Parameters
For the configuration there are only two parameters to be set:
- Init. grid size - in general it can be seen as an average superpixels size.
- Regularisation - influence the compactness of estimated superpixels. The range is from 0 (very elastic superpixels) to 1 (superpixels are nearly squares). Experimentally, we set as optimal value 0.2 for most cases.
Visualisation
To show of handle segmentation results we presented a few approaches:
- Overlap contours - simply draw the contours on resulting superpixels into the image by chosen colour.
- Export segments as ROIs - all superpixels are exported as polygons into the ROI Manager.
- Show final segmentation - add one more layer and fill each superpixel by a random colour.
- Save segmentation into file - export the superpixel segmentation into a text file as segmentation matrix with labels. The first line mark the image dimensions (Dims: {width} {height}) and then follow the labeling where each number represents the superpixel index. (Note, number of lines is equal to {image width} and there is {image height} number of indexes which are splitted by blanc space.) Have a look at sample file
.
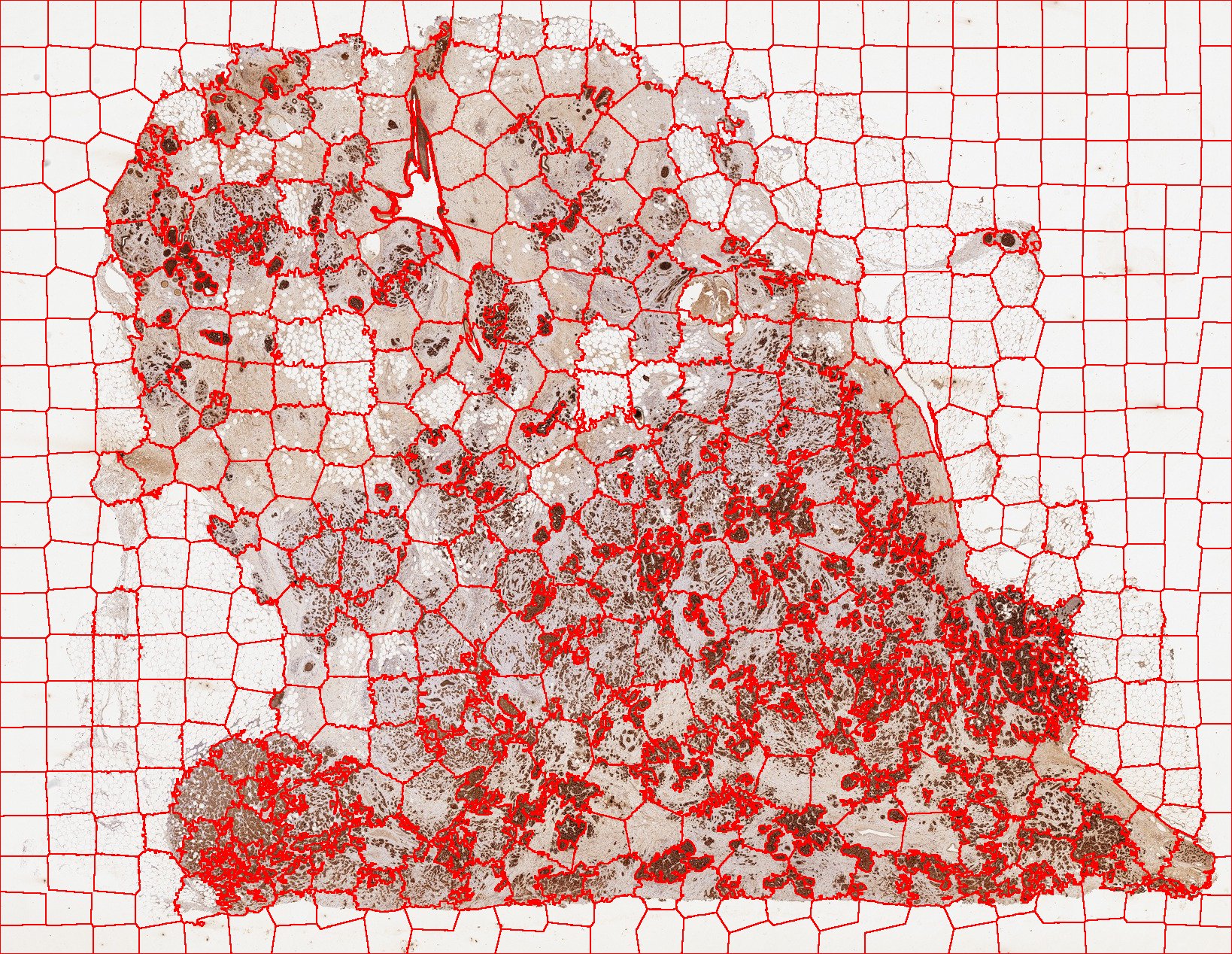
Sample of jSLIC segmented histological tissue
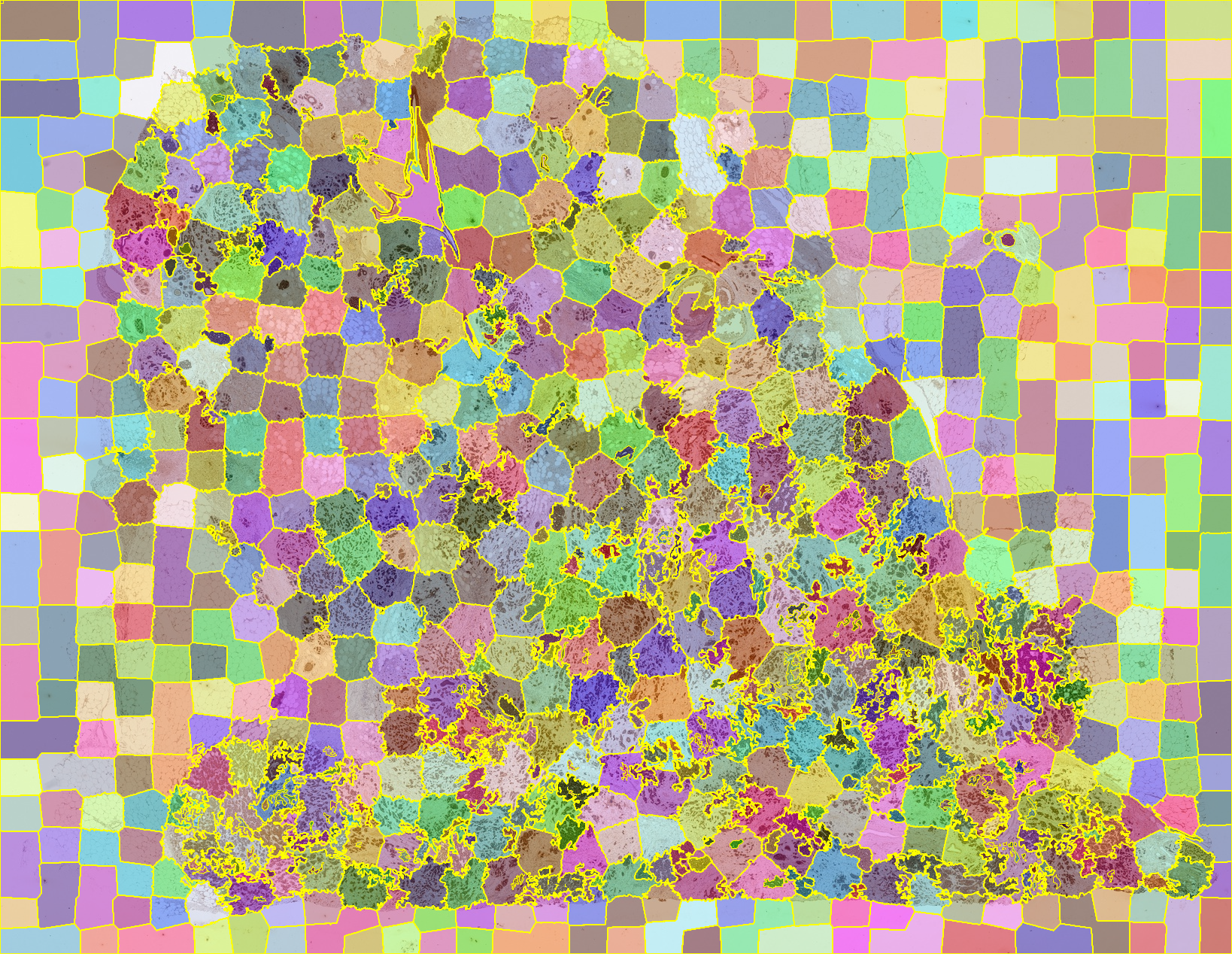

Sample of jSLIC segmented histological tissue
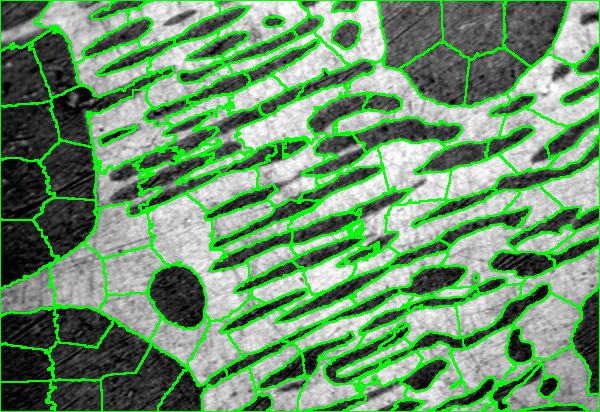
Sample of jSLIC segmented AuPbSn40
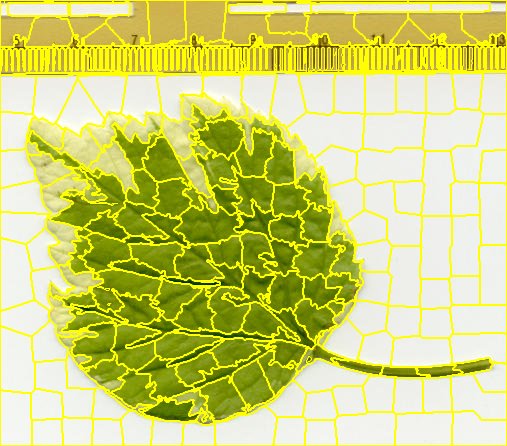
Sample of jSLIC segmented Leaf
References
Borovec, J., & Kybic, J. (2014). jSLIC : superpixels in ImageJ. Computer Vision Winter Workshop. Praha.